Descriptors¶
Introduction¶
The ccdc.descriptors module contains functionality for generating
geometric, molecular and crystallographic descriptors.
Let us create an ccdc.io.EntryReader so that we can read in
molecules and crystals from the CSD.
>>> from ccdc.io import EntryReader
>>> entry_reader = EntryReader('CSD')
As a preamble let us also set up a variable for a temporary directory.
>>> import tempfile
>>> tempdir = tempfile.mkdtemp()
Molecular geometry¶
The module ccdc.descriptors.MolecularDescriptors contains a number of static methods for inspecting the geometry of a molecule. For example,
>>> from ccdc.descriptors import MolecularDescriptors as MD, GeometricDescriptors as GD
>>> aabhtz = entry_reader.molecule('AABHTZ')
>>> print(round(MD.atom_distance(aabhtz.atom('Cl1'), aabhtz.atom('Cl2')), 2))
5.47
>>> print(round(MD.atom_angle(aabhtz.atom('Cl1'), aabhtz.atom('C6'), aabhtz.atom('Cl2')), 2))
178.25
>>> print(round(MD.atom_torsion_angle(aabhtz.atom('Cl1'), aabhtz.atom('Cl2'), aabhtz.atom('O2'), aabhtz.atom('O1')), 2))
12.45
Note that the atoms do not have to be connected for the geometry to be determined.
One can construct centroids of sets of atoms, vectors from two atoms and RMSD fitted planes. For convenience one can construct centroids and planes directly from a ring:
>>> centroid = MD.atom_centroid(*tuple(a for a in aabhtz.atoms))
>>> r1_centroid = MD.ring_centroid(aabhtz.rings[0])
>>> r2_centroid = MD.ring_centroid(aabhtz.rings[1])
>>> r1_plane = MD.ring_plane(aabhtz.rings[0])
>>> r2_plane = MD.ring_plane(aabhtz.rings[1])
The plane methods construct instances of ccdc.descriptors.GeometricDescriptors.Plane. This object supports a few methods for geometric analysis as well as the normal vector and distance from the origin:
>>> print(round(GD.point_distance(r1_centroid, r2_centroid), 3))
6.385
>>> print(round(r1_plane.plane_angle(r2_plane), 3))
88.576
>>> print(round(r1_plane.point_distance(r1_centroid), 3))
0.0
>>> print(r1_plane.normal)
Vector(0.608, -0.050, 0.792)
>>> print(round(r1_plane.distance, 3))
-1.785
Vectors and planes may be constructed directly from points:
>>> x_axis = GD.Vector(1, 0, 0)
>>> vec = GD.Vector.from_points(r1_centroid, centroid)
>>> plane = GD.Plane.from_points(centroid, r1_centroid, r2_centroid)
With the module ccdc.descriptors.MolecularDescriptors is also possible to calculate the adjacency matrix of a molecule. For instance, the following code allows one to calculate the logarithm of the number of walks of length k that start and end at the same atom.
>>> abeboz = entry_reader.molecule('ABEBOZ')
>>> adjacency_matrix = MD.AdjacencyMatrixDescriptorCalculator(abeboz)
>>> print(round(adjacency_matrix.self_returning_walk_ln(10), 3))
9.838
It is possible to count the number of times a pair of elements appear with a specified minimum path length. For example:
>>> atom_pair_distance = MD.AtomPairDistanceDescriptorCalculator(abeboz)
>>> print(atom_pair_distance.element_pair_count("C","N",5))
2.0
Finally, one can look at the connectivity of a molecule and calculate a connectivity index:
>>> connectivity_index = MD.ConnectivityIndices(abeboz)
>>> print(round(connectivity_index.connectivity_index(4), 3))
5.861
Superimposing molecules¶
Suppose that we want to superimpose a set of conformers using a substructure of the input molecule rather than all the atoms in it.
This can be achieved using the ccdc.descriptors.MolecularDescriptors.Overlay.
Let us generate a set of 25 conformers of Rosuvastatin using the PDB ligand from 1hwl as the starting conformation.
>>> from ccdc.io import MoleculeReader
>>> filepath = '1hwl-lig.mol2'
In this instance we will get the first molecule in the file.
>>> mol_reader = MoleculeReader(filepath)
>>> mol = mol_reader[0]
>>> mol_reader.close()
Now we set up a conformer generator and generate the 25 conformations.
>>> from ccdc.conformer import ConformerGenerator
>>> conformer_generator = ConformerGenerator()
>>> conformer_generator.settings.max_conformers = 25
>>> conformers = conformer_generator.generate(mol)
In this instance we want to superimpose the conformers on the pyrimidine ring of the input molecule. We therefore generate a pyrimidine substructure search.
>>> from ccdc.search import SubstructureSearch, SMARTSSubstructure
>>> searcher = SubstructureSearch()
>>> sub_id = searcher.add_substructure( SMARTSSubstructure('c1ncncc1') )
We can use this to identify the atoms to match in the input molecule.
>>> input_hits = searcher.search(mol)
>>> print(len(input_hits))
1
>>> input_match_atoms = input_hits[0].match_atoms()
Finally, we can use this information to write out a file with conformers
superimposed on the pyridine ring of the input structure using the
ccdc.descriptors.MolecularDescriptors.Overlay.
>>> from ccdc.descriptors import MolecularDescriptors
>>> from ccdc.io import MoleculeWriter
>>> mol_writer = MoleculeWriter(os.path.join(tempdir, 'superpos.mol2'))
>>> for c in conformers:
... cmol = c.molecule
... chits = searcher.search(cmol)
... match_atoms = chits[0].match_atoms()
... atom_pairs = zip(input_match_atoms, match_atoms)
... overlay = MolecularDescriptors.Overlay(mol, cmol, atoms=atom_pairs)
... overlaid_mol = overlay.molecule
... mol_writer.write(overlaid_mol)
...
Here overlay.molecule returns input molecule cmol transformed to overlay onto mol Other useful properties of the overlayed results are: : rmsd - Returns RMSD between the two overlaid molecules : rmsd_tanimoto - Returns Tanimoto RMSD between the two overlaid molecules : transformation - Returns Molecule.Transformation object required to overlay mol2 over mol1 : max_distance - Returns the maximum distance between two equivalent atoms in the overlay (Angstroms).
>>> from ccdc.descriptors import MolecularDescriptors
>>> mol1 = entry_reader.molecule('NALCYS02')
>>> mol2 = entry_reader.molecule('NALCYS17')
>>> overlay = MolecularDescriptors.Overlay(mol1, mol2)
>>> molecule = overlay.molecule
>>> rmsd = overlay.rmsd
>>> rmsd_tanimoto = overlay.rmsd_tanimoto
>>> trans = overlay.transformation
>>> max_distance = overlay.max_distance
>>> print(f"{mol1.identifier} v {mol2.identifier} :")
NALCYS02 v NALCYS17 :
>>> print(f"RMSD : {rmsd:.3f} A")
RMSD : 0.999 A
>>> print(f"Max Distance : {max_distance:.3f} A")
Max Distance : 2.228 A
>>> print(f"Tanimoto RMSD : {rmsd_tanimoto:.3f} A")
Tanimoto RMSD : 0.816 A
Another function of interest in this context is the
ccdc.descriptors.MolecularDescriptors.rmsd().
Maximum Common Substructure¶
ccdc.descriptors.MolecularDescriptors.MaximumCommonSubstructure may be used to find a maximum common substructure of two ccdc.molecule.Molecule instances. We can create an instance of the appropriate class:
>>> mcss = MolecularDescriptors.MaximumCommonSubstructure()
This instance contains a nested class, ccdc.descriptors.MolecularDescriptors.MaximumCommonSubstructure.Settings allowing the configuration of graph search. The meaning of the properties should be evident from their names. Their default settings are as follows:
>>> print(mcss.settings.ignore_hydrogens)
False
>>> print(mcss.settings.check_bond_count)
False
>>> print(mcss.settings.check_bond_polymeric)
False
>>> print(mcss.settings.check_bond_type)
True
>>> print(mcss.settings.check_charge)
True
>>> print(mcss.settings.check_element)
True
>>> print(mcss.settings.check_hydrogen_count)
False
Let us read in two molecules with a non-trivial common substructure:
>>> csd = MoleculeReader('CSD')
>>> hxacan = csd.molecule('HXACAN')
>>> wexwut = csd.molecule('WEXWUT')
We can now determine the maximum common substructure of these two molecules. With default settings only the phenyl ring is common, so we will relax the conditions of the search:
>>> mcss.settings.check_element = False
>>> mcss.settings.ignore_hydrogens = True
>>> atoms, bonds = mcss.search(hxacan, wexwut)
The atoms returned is a tuple of pairs of atoms matched, the first from HXACAN, the second from WEXWUT. Similarly for matched bonds. We can extract the atoms for either molecule using python’s zip function:
>>> hxacan_atoms = list(zip(*atoms))[0]
>>> wexwut_atoms = list(zip(*atoms))[1]
For example, let us make a diagram, highlighting the common substructure in each molecule. Now we can see that with the relaxed search conditions all of HXACAN is matched with a similar group in WEXWUT:
>>> from ccdc.diagram import DiagramGenerator
>>> generator = DiagramGenerator()
>>> generator.settings.highlight_color = 'cyan'
>>> generator.settings.shrink_symbols = False
>>> hxacan_image = generator.image(hxacan, highlight_atoms=hxacan_atoms)
>>> wexwut_image = generator.image(wexwut, highlight_atoms=wexwut_atoms)
>>> hxacan_image.save('hxacan_mcss.png')
>>> wexwut_image.save('wexwut_mcss.png')
HXACAN |
WEXWUT |
|---|---|
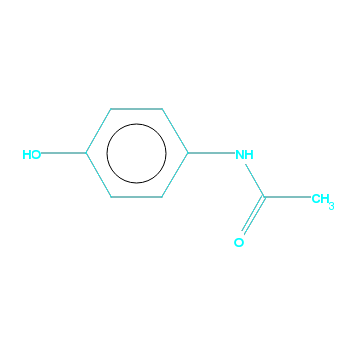
|

|
There is a class, ccdc/descriptors.MolecularDescriptors.AtomDistanceSearch which allows rapid proximity detection for atoms. This is of most use when dealing with proteins, but can be used for small molecule proximity detection. For example to look for possible close contacts:
>>> searcher = MD.AtomDistanceSearch(aabhtz)
>>> for a in aabhtz.heavy_atoms:
... contacts = set(searcher.atoms_within_range(a.coordinates, 2.0))
... contacts = contacts - set(a.neighbours) - set([a])
... if contacts:
... print(a, contacts)
Atom(N1) set([Atom(H4)])
Atom(N5) set([Atom(H6)])
Atom(C10) set([Atom(H9), Atom(H6), Atom(H7)])
See also
For more information on the available molecular descriptors
please have a look at the API documentation:
ccdc.descriptors.MolecularDescriptors.
Pore Analyser¶
To demonstrate use of the ccdc.descriptors.CrystalDescriptors.PoreAnalyser class we can
read in a crystal from the CSD (although a CIF file would work equally well).
>>> crystal = entry_reader.crystal('AKUKUO')
To use the Pore Analyser tool, we need to add the structure of interest to the analyser
>>> from ccdc.descriptors import CrystalDescriptors
>>> pore_analyser = CrystalDescriptors.PoreAnalyser(crystal)
Each value of interest can then be obtained by calling the property, for example the total surface area:
>>> print(round(pore_analyser.total_surface_area,2))
58.51
Settings can be used to control the various parameters for the pore analyser calculation.
>>> settings = CrystalDescriptors.PoreAnalyser.Settings()
>>> settings.n_probe_sigma = 2.0
>>> pore_analyser = CrystalDescriptors.PoreAnalyser(crystal, settings)
>>> print(round(pore_analyser.total_surface_area, 2))
165.07
Powder patterns¶
Note
The powder pattern features are available only to CSD-Materials and CSD-Enterprise users.
Simulating powder patterns¶
To illustrate the use of the ccdc.descriptors.CrystalDescriptors.PowderPattern
class let us read in a crystal from the CSD.
>>> crystal = entry_reader.crystal('AFUSEZ')
To create a powder pattern we import the
ccdc.descriptors.CrystalDescriptors.PowderPattern class and use its
ccdc.descriptors.CrystalDescriptors.PowderPattern.from_crystal() static method.
>>> from ccdc.descriptors import CrystalDescriptors
>>> pattern = CrystalDescriptors.PowderPattern.from_crystal(crystal)
The pattern created is an instance of the
ccdc.descriptors.CrystalDescriptors.PowderPattern class.
To write the pattern to as a xye or Bruker raw file one can use the
functions ccdc.descriptors.CrystalDescriptors.PowderPattern.write_xye_file() and
ccdc.descriptors.CrystalDescriptors.PowderPattern.write_raw_file() respectively.
>>> pattern.write_xye_file(os.path.join(tempdir, 'afusez.xye'))
Comparing powder patterns¶
It is possible to read in a pattern stored in xye format using
the ccdc.descriptors.CrystalDescriptors.PowderPattern.from_xye_file() function.
>>> file_pattern = CrystalDescriptors.PowderPattern.from_xye_file(os.path.join(tempdir, 'afusez.xye'))
Let us determine the similarity between the AFUSEZ and AFUSEA crystals’ calculated powder patterns.
>>> afusea_pattern = CrystalDescriptors.PowderPattern.from_crystal(entry_reader.crystal('AFUSEA'))
>>> round( file_pattern.similarity(afusea_pattern), 3 )
0.92
Settings can be used to control the various parameters of the simulation of a powder pattern. For example to use a non-default wavelength and non-standard two theta range:
>>> settings = CrystalDescriptors.PowderPattern.Settings()
>>> settings.wavelength = CrystalDescriptors.PowderPattern.Wavelength(CrystalDescriptors.PowderPattern.Wavelength.Wavelength_FeKa1)
>>> settings.two_theta_minimum = 10.
>>> settings.two_theta_maximum = 70.
>>> pattern = CrystalDescriptors.PowderPattern.from_crystal(crystal, settings)
See also
Statistical descriptors¶
There is a namespace, ccdc.descriptors.StatisticalDescriptors for statistical measures
which we have found useful. There is no intent to make this a complete statistical package such
as Rpy but it has been found convenient to have implementations
of enrichment metrics when analysing results from virtual screening and docking studies.
Within this namespace is a class, ccdc.descriptors.StatisticalDescriptors.RankStatistics
for calculating various rank-based enrichment metrics.
Firstly let us import the appropriate module:
>>> import operator
>>> import collections
>>> from ccdc.descriptors import StatisticalDescriptors
Next we will set up some dummy activity data.
>>> activity_data_file = 'rank_statistics.csv'
We will now read the file and format the appropriate data:
>>> with open(activity_data_file) as f:
... rows = [l.split(',') for l in f]
>>> scores = [(float(row[0].strip()), bool(int(row[1].strip())), row[2]) for row in rows]
The ccdc.descriptors.StatisticalDescriptors.RankStatistics may now be initialised with these data:
>>> rank_stats = StatisticalDescriptors.RankStatistics(scores, activity_column=operator.itemgetter(1))
The argument, activity_column indicates which column of the scores data should be interpreted as
an activity classification. In this instance we have used operator.itemgetter. We can instead
use operator.attrgettr to extract activity data from class instances. An artificial example is
given below:
>>> ActivityRecord = collections.namedtuple('ActivityRecord', ['score', 'active', 'identifier'])
>>> activity_records = [ActivityRecord(*row) for row in scores]
>>> rank_stats = StatisticalDescriptors.RankStatistics(scores, activity_column=operator.attrgetter('activity'))
For convenience, if the scores data is an indexed collection, as it is in this example, then a simple integer index may be used to specify the column with the activity classification. Similarly, the activity column may be specified as a string field name for activity data in namedtuples.
>>> rank_stats = StatisticalDescriptors.RankStatistics(scores, activity_column=1)
When the scores are an iterable of booleans the activity_column need not be specified:
>>> activities = [row[1] for row in scores]
>>> rank_stats = StatisticalDescriptors.RankStatistics(activities)
The methods of the ccdc.descriptors.StatisticalDescriptors.RankStatistics now return the
various enrichment metrics:
>>> print(round(rank_stats.AUC(), 3))
0.725
>>> print(round(rank_stats.EF(0.1), 3))
4.468
>>> print(round(rank_stats.ACC(0.1), 3))
0.897
>>> print(round(rank_stats.PPV(0.1), 3))
0.099
>>> print(round(rank_stats.EF(0.5), 3))
1.362
>>> print(round(rank_stats.ACC(0.5), 3))
0.507
>>> print(round(rank_stats.PPV(0.5), 3))
0.03
Principle axes aligned molecules¶
There is a class, ccdc.descriptors.MolecularDescriptors.PrincipleAxesAlignedBox which allows aligning of a molecule on its principle axes, and deriving a minimal bounding box around it.
Let us take an example, paracetamol, from the CSD and explore its packing shell to see how it grows as hydrogen bonds are expanded.
>>> hxacan_crystal = entry_reader.crystal('HXACAN')
>>> hxacan_mol = hxacan_crystal.molecule
We can now form its minimal bounding box:
>>> box = MolecularDescriptors.PrincipleAxesAlignedBox(hxacan_mol)
The primary, secondary and tertiary axes, and the volume of the box are accessible as properties:
>>> print('%s, %s, %s' % (box.x_vector, box.y_vector, box.z_vector))
Vector(7.448, 5.724, 6.326), Vector(4.769, 3.794, 2.816), Vector(3.264, 1.785, 2.004)
>>> print(round(box.volume, 3))
321.265
The other property of import is the aligned molecule:
>>> aligned_mol = box.aligned_molecule
Ligand Overlay¶
The ligand overlay tool is available to use in the CSD Python API as of version 3.4.0. In this example we will overlay two antihistamines: Ipratroponium and Tiotroponium. These molecules share similar features and are shown below in a matching orientation, as they would appear in a protein cavity.
Ipratroponium |
Tiotroponium |
|---|---|

|

|
The ligand overlay process has not been significantly altered from the GUI version; it has simply been exposed in the Python API. As a result, all the original files are still created, and the algorithm itself will produce the same overlays as before.
There is a new class ccdc.descriptors.MolecularDescriptors.LigandOverlay. Ligands also need to be supplied to this class through the usual ccdc.io.MoleculeReader class, commonly used throughout our API.
>>> from ccdc.descriptors import MolecularDescriptors
>>> from ccdc.io import MoleculeReader
This class takes a single instantiation argument: a string defining the location to run the overlay from.
>>> overlay = MolecularDescriptors.LigandOverlay('.')
>>> ipratroponium_file = 'ipratroponium.mol2'
>>> tiotroponium_file = 'tiotroponium.mol2'
ccdc.io.MoleculeReader is quite flexible in the files it can read, including accepting multiple molecules at once through an SDF. Having read molecules through the MoleculeReader class, they may be supplied through the add_ligands() method. This method simply takes a list of molecules, here we supply only two, but more may be added.
>>> mol1 = MoleculeReader(ipratroponium_file)[0]
>>> mol2 = MoleculeReader(tiotroponium_file)[0]
>>> overlay.add_ligands([mol1, mol2])
Finally, the ligand overlay process is started with the overlay_ligands() method. This will run the ligand overlay, and generate all the usual files. Your terminal should show the progress of the overlay tool as it runs.
>>> results = overlay.overlay_ligands()
By assigning this call to a variable, we can inspect the results.
>>> for result in results:
... print(result.hbond)
... print(result.volume)
... print(result.hydrophobe)
16.0
...
The output files will contain a series of SDF and mol2 files. These are the overlaid results which can be viewed through Hermes, for example.
